What Stimulates The Ribosome To Move Down One Codon
So, imagine this: I’m staring at a ridiculously complicated diagram of a cell. You know, the kind that makes you feel like you need a PhD just to identify a mitochondrion. My eyes land on this little, whirring contraption called a ribosome, diligently churning out proteins. It looks like a microscopic factory, and I’m thinking, “Okay, so it’s reading instructions, but what’s actually pushing it along? What’s the tiny engine that makes it chug down the mRNA chain, codon by codon?” It’s like wondering what makes a train move down the track – is it the engine, the wheels, the driver? Or is it something more subtle, like the magnetic pull of the rails?
This whole process, this incredible dance of molecules, has always fascinated me. It’s not just about making proteins, right? It’s about life itself. And the ribosome, this protein-making machine, is at the heart of it all. It’s where the genetic code, the blueprint of our existence, is translated into the functional machinery of our cells.
But how does it move? I mean, it’s not like it has tiny little legs or a built-in conveyor belt system. It's an intricate molecular ballet, and the conductor of this ballet, the one that cues the next step, is what we’re going to dive into. Because honestly, it’s way cooler than any sci-fi movie you’ve seen.
The Ribosome: A Protein-Making Marvel
Before we get to the nitty-gritty of movement, let’s just take a moment to appreciate the ribosome. These things are absolutely ubiquitous. Every single living cell, from the tiniest bacterium to the most complex human, has ribosomes. They are the universal builders, the tireless workers of the cellular world.
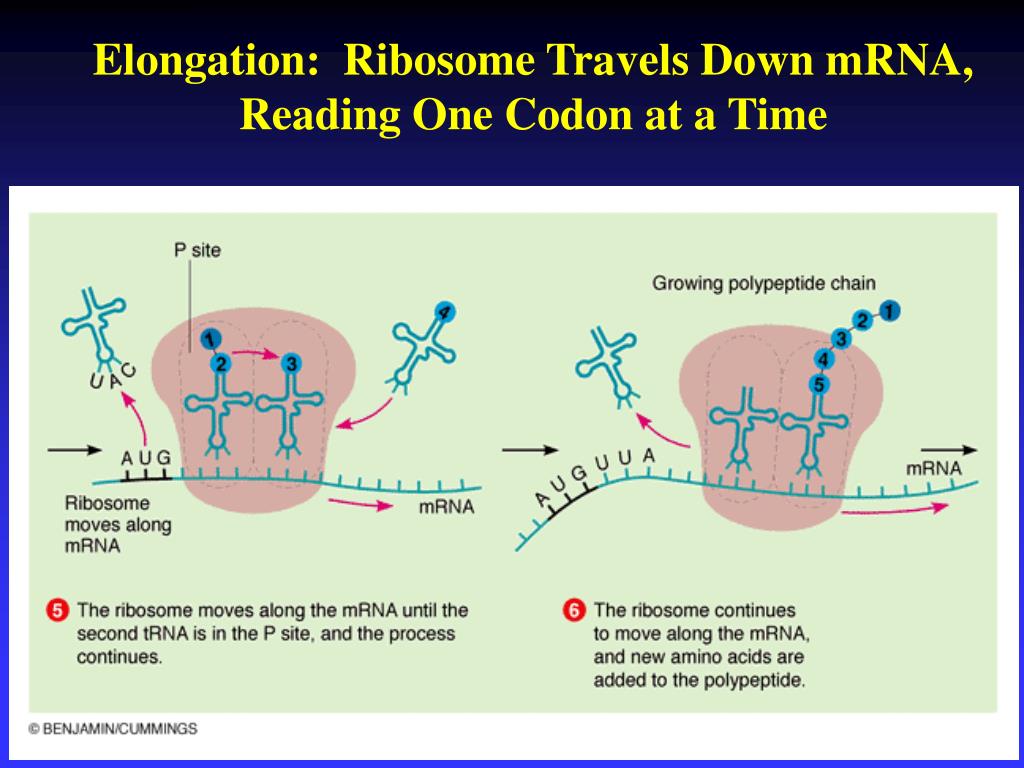
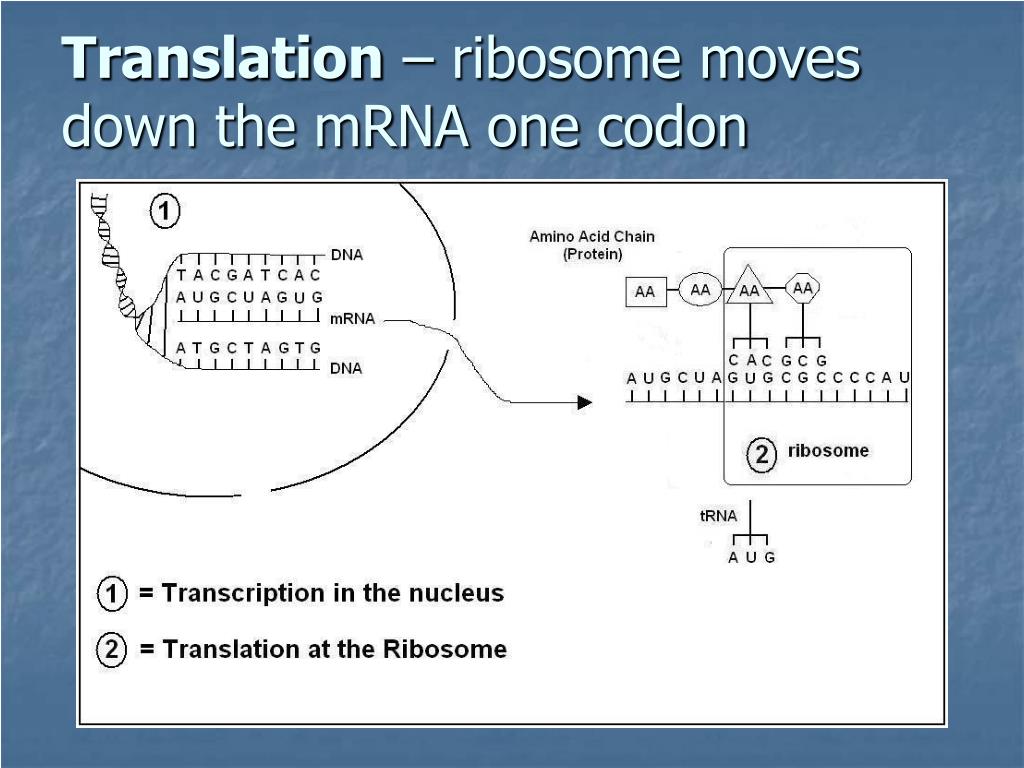
Think of the mRNA as a long, single-stranded ribbon carrying the genetic instructions from the DNA. The ribosome latches onto this ribbon and starts reading it, three bases at a time. These three-base sequences are called codons, and each codon specifies a particular amino acid. The ribosome then recruits the correct amino acid, carried by another molecule called tRNA (transfer RNA), and links them together to form a growing chain of amino acids – that's your protein!
It's a remarkably accurate process. We’re talking about a precision that would make a Swiss watchmaker weep with joy. But accuracy isn't the only thing that matters. For this process to work, the ribosome needs to move. It can’t just sit there reading the same codon forever. It needs to advance, one codon at a time, to build the entire protein sequence.
The "Codon Hop": What Exactly Makes It Move?
Okay, so the big question: what stimulates this movement? It’s not a single, dramatic event, like a giant hand pushing the ribosome along. Instead, it’s a beautiful interplay of chemical signals and molecular shape changes. And the star of the show, the molecule that’s really pulling the strings (or should I say, the codons?), is GTP.
You might have heard of ATP (adenosine triphosphate), the main energy currency of the cell. GTP (guanosine triphosphate) is its close cousin. They’re very similar in structure and function, and in the case of protein synthesis, GTP is the key player in ribosome translocation, which is the fancy term for the ribosome’s movement down the mRNA.

So, how does it work? Let’s break it down. At any given moment, the ribosome has three distinct sites where tRNA molecules can bind: the A site (aminoacyl site), the P site (peptidyl site), and the E site (exit site).
When the ribosome first starts, it’s usually poised at the start codon on the mRNA. A tRNA carrying the first amino acid (methionine, in most cases) will bind to the P site.
Now, here’s where the magic starts. For the ribosome to move, it needs to shift its position on the mRNA. This involves a complex series of conformational changes within the ribosome itself, orchestrated by a special protein called elongation factor G (EF-G) in bacteria, or eukaryotic elongation factor 2 (eEF2) in eukaryotes. These elongation factors are like the skilled mechanics of the ribosome, ensuring smooth operation.
EF-G/eEF2 binds to the ribosome when it’s in a particular state, and here’s the crucial part: it needs GTP to do its job. It hydrolyzes GTP (breaks it down into GDP and inorganic phosphate), releasing energy. This energy fuels a conformational change in EF-G/eEF2 itself.
Imagine EF-G/eEF2 as a spring-loaded mechanism. When it binds GTP, it’s in a relaxed state. When it hydrolyzes GTP, it becomes “cocked” or energized. This energized state allows it to interact with the ribosome in a way that forces the ribosome to move along the mRNA by one codon.
Think of it like this: the elongation factor, powered by GTP, essentially “pushes” the ribosome. It slides the ribosome forward, precisely one codon at a time. This movement is what shifts the tRNA molecules within the ribosome. The tRNA that was in the A site (carrying the newly added amino acid) moves to the P site. The tRNA that was in the P site (carrying the growing polypeptide chain) moves to the E site, where it will be released.
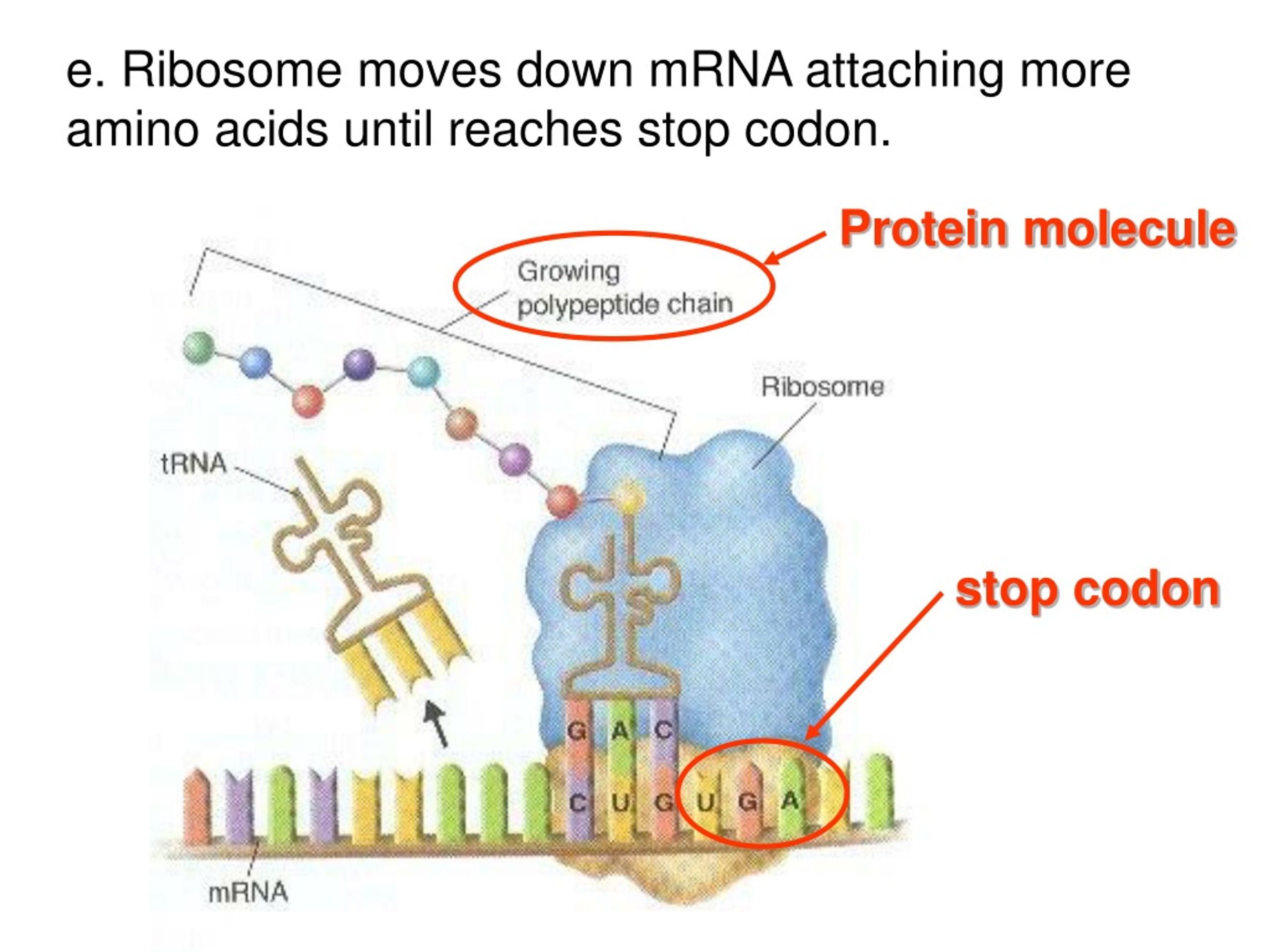
After the ribosome has moved, the EF-G/eEF2, now in its GDP-bound form, detaches from the ribosome. It’s like the mechanic finishing their job and moving on. The ribosome is now ready for the next cycle of elongation: a new charged tRNA enters the now-empty A site, and the process repeats.
The Role of GTP Hydrolysis: The Cellular Engine
So, the hydrolysis of GTP is absolutely fundamental. It's not just about providing energy; it's about the conformational changes it enables. Without GTP, EF-G/eEF2 wouldn’t be able to undergo the structural rearrangements necessary to force the ribosome to translocate.
This is a common theme in molecular biology. Many cellular processes rely on the binding and hydrolysis of GTP to regulate protein function and drive mechanical movements. GTP-binding proteins, or G proteins, act like molecular switches. They are active when bound to GTP and inactive when bound to GDP.
In the case of ribosome movement, EF-G/eEF2 acts as a G protein. It cycles between a GTP-bound, active state (where it can promote translocation) and a GDP-bound, inactive state (where it detaches and waits for another GTP to bind). This constant cycling ensures that the ribosome moves in a controlled and unidirectional manner.
It's quite elegant, isn't it? This tiny molecule, GTP, is the fuel that powers the movement of this incredibly complex molecular machine. It’s a testament to the efficiency and sophistication of cellular processes.
Beyond Elongation Factors: Other Nuances
While EF-G/eEF2 and GTP are the primary drivers of ribosome translocation, it's worth mentioning that the ribosome itself isn't a static structure. It's a dynamic entity with many moving parts. The different ribosomal RNA (rRNA) molecules and ribosomal proteins within the ribosome undergo subtle shape changes during each step of translation.
These internal movements, coupled with the external push from EF-G/eEF2, ensure that the mRNA moves through the ribosome in a precise, step-by-step fashion. It’s not just a simple sliding; it's a more complex "ratcheting" motion.
Think of a ratchet mechanism. It allows movement in one direction but prevents backward motion. This ensures that the ribosome always moves forward, codon by codon, without accidentally slipping back. This directional control is absolutely critical for accurate protein synthesis.
Also, there are other factors involved, especially in eukaryotes. For instance, the process of "release" of the polypeptide chain involves different elongation factors and GTP, but the fundamental principle of GTP-driven molecular machinery remains the same.
The "Mistake" Scenario: When Things Go Wrong
You might be wondering, what happens if the ribosome reads the "wrong" codon, or if a tRNA with the wrong amino acid tries to bind? Well, there are proofreading mechanisms involved, but sometimes mistakes happen. However, the focus of this article is on the forward movement. The translocation step, driven by EF-G/eEF2 and GTP, is generally a very reliable process.
Interestingly, some toxins and antibiotics work by interfering with this process. For example, certain antibiotics can bind to the ribosome and block the action of EF-G/eEF2 or prevent GTP hydrolysis, thereby halting protein synthesis. This is why these antibiotics are so effective against bacteria – they shut down the essential protein-making machinery of the enemy.

It's quite a dangerous business for a bacterium to have its ribosomes messed with! Imagine your own cellular factory grinding to a halt. Not ideal for survival.
A Cosmic Choreography
Looking at the whole picture, it’s mind-boggling. We have this intricate system where genetic information is transcribed into mRNA, then translated by ribosomes, with the help of tRNA and powered by GTP. Each step is perfectly timed and coordinated.
The ribosome moves because a specific protein, EF-G/eEF2, bound to GTP, undergoes a shape change after hydrolyzing the GTP. This shape change physically pushes the ribosome along the mRNA strand by one codon. It’s a controlled, energy-dependent movement that ensures the sequential addition of amino acids to the growing polypeptide chain.
It's a bit like a dancer performing a complex routine. The music (mRNA sequence) dictates the steps, the choreographer (EF-G/eEF2) guides the dancer (ribosome), and the dancer's own energy and movements (GTP hydrolysis and conformational changes) make the routine possible. And the result? A beautiful, functional protein, ready to do its job in the cell.
So, the next time you think about what makes life tick, remember the humble ribosome and its GTP-fueled journey down the mRNA. It's a testament to the power of molecular machinery and the elegance of biological design. It’s not just moving; it’s building the very fabric of life, one codon at a time. And that, my friends, is pretty darn amazing if you ask me. We're talking about the fundamental engine of life itself, right there in your cells, humming away. Pretty wild, huh?
It's a constant reminder that even the smallest, most fundamental processes in our bodies are incredibly complex and beautifully orchestrated. The more you look, the more you realize there’s an entire universe of intricate mechanisms at play, all working in harmony. And it all starts with reading that genetic code, codon by codon, propelled by the subtle yet powerful force of GTP.
