What Is The First Step In A Pulse Chase Experiment
You know, I remember this one time, ages ago, when I was first dipping my toes into the thrilling, and sometimes utterly baffling, world of molecular biology. I was fiddling with some kind of Western blot, convinced I was on the cusp of a Nobel-winning discovery (spoiler alert: I wasn't). Anyway, I’d spent hours, literal hours, meticulously pipetting, incubating, washing… you name it. My lab bench looked like a crime scene, but instead of blood, it was filled with little plastic tubes and colourful liquids. And then, just as I was about to declare victory, I realized I’d completely forgotten to add the actual thing I was supposed to be detecting. Like, the main ingredient. My whole experiment was… well, it was a very expensive, very complicated way to prove that nothing happened. Cue the dramatic sigh.
It’s funny, isn't it? How one tiny, seemingly insignificant step, or the complete lack of it, can render your entire meticulous effort utterly useless? That’s kind of what got me thinking about pulse-chase experiments. If you've ever stumbled across this fancy-sounding technique, you might imagine it involves some kind of daring, high-speed pursuit of molecules. And in a way, you wouldn't be entirely wrong!
The "Pulse" Part: Getting Their Attention
So, what is a pulse-chase experiment, anyway? At its heart, it's a way to track the journey of molecules within a cell or organism over time. Think of it like this: you want to know where your kid’s favourite toy goes after they’ve finished playing with it. Is it under the sofa? Did the dog abscond with it? Or did it magically teleport to Narnia? A pulse-chase experiment is the scientific equivalent of putting a tiny, invisible GPS tracker on that toy. Pretty neat, huh?
Now, the name itself gives us a pretty big clue. We've got the "pulse" and the "chase." Let's tackle the "pulse" first. This is where the whole "getting their attention" thing comes in. Imagine our molecules are a whole bunch of shy introverts at a party. They're there, mingling, but you can't really tell who’s who, or which ones are doing what. The "pulse" is like a sudden, bright spotlight that you shine on a specific group of these molecules.
How do we do that? Well, it usually involves introducing a labelled molecule into the system. This label is like a little flag, a beacon, that allows us to specifically detect and track those molecules later on. What kind of labels? Oh, the possibilities are endless! We could use radioactive isotopes, like 3H or 14C, which are like little glowing tags that our detectors can pick up. Or, if we're feeling a bit more modern and perhaps a tad less radioactive (which is often a good thing for everyone involved!), we might use fluorescent labels. These are like tiny, colourful LEDs that we can see under a special microscope. They’re so cool, seriously.
The "pulse" itself is a short, defined period where we expose our cells or organism to this labelled molecule. It's like a brief flash of that spotlight. We're not giving them the label for hours and hours. Oh no. That would be like keeping the spotlight on the entire party, making everyone equally visible. We want to label a specific cohort, a generation, of molecules. So, we give them a taste of the labelled stuff for a short burst. Think minutes, or maybe a couple of hours, depending on what we’re studying.
The Crucial First Step: What Exactly Is That Pulse?
So, if the pulse is this crucial introductory phase, what is the very first thing you absolutely must get right? Drumroll, please… it’s the selection and introduction of the labelled precursor molecule. Yep, it sounds simple, almost anticlimactic, doesn't it? But trust me, this is the bedrock upon which your entire pulse-chase experiment is built. If you pick the wrong precursor, or introduce it incorrectly, your entire elaborate chase is going to end up like my blot experiment – a lot of effort, zero results.
Let’s break this down a bit, because there’s more to it than just grabbing a bottle. You’re not just grabbing any labelled molecule. You have to choose one that is specifically incorporated into the molecule you’re interested in. For example, if you want to track newly synthesized proteins, you’d typically use a labelled amino acid, like methionine or leucine. The cell, bless its little heart, will happily grab that labelled amino acid and use it as a building block to churn out new proteins. Bingo!
Or, if you’re interested in DNA replication, you might use a labelled nucleotide, like thymidine. The cell, busy copying its genetic material, will incorporate that labelled nucleotide into the new DNA strands. Ingenious, right?
The key here is that the labelled precursor has to be something the cell naturally uses. You're not forcing it to do anything unnatural; you're just giving it the building blocks with a little "I was here!" sign attached. This specificity is absolutely paramount. If you accidentally feed it something that gets incorporated into multiple different molecules, you’ll end up with a jumbled mess, and distinguishing your target molecule will be like trying to find a specific grain of sand on a beach – impossible.
And it’s not just about what you choose, but also how you introduce it. Are you adding it to the cell culture medium? Injecting it? This depends entirely on your experimental system. The goal is to ensure efficient uptake and incorporation by the target cells or organism. You want a strong, clear pulse, not a faint whisper that gets lost in the noise.
Think about it like this: you’re trying to get everyone in a crowded room to wear a specific coloured hat for a few minutes. If you just throw a pile of hats in the middle, some people might grab one, some might not, and it’ll be a chaotic free-for-all. But if you hand out the hats directly to each person, ensuring they put one on, you get a much cleaner, more controlled effect. That’s what good precursor introduction is all about.
So, before you even think about waiting for the "chase" part to begin, before you plan your sampling times, before you get excited about your fancy fluorescent microscopes, you need to spend some serious time thinking about that labelled precursor. Is it the right one? Will it be incorporated? Will it give you a strong enough signal? This is the foundation. Get this wrong, and the whole edifice crumbles.
The "Chase": Following the Trail
Okay, so you've successfully pulsed your molecules. They've got their little flags, their little beacons. They're all marked, ready to be followed. Now comes the "chase." This is where the real detective work begins. After the initial, short pulse of the labelled precursor, you remove it from the environment. You wash it all away, you give them fresh, unlabelled building blocks, or you move them to an environment where the labelled precursor is no longer available.
This is crucial. It's like taking away the bag of special coloured hats. Now, everyone who has a hat will continue wearing it, but any new hats that are made will be the plain, unlabelled kind. This ensures that at any given time point after the pulse, the radioactivity or fluorescence you detect will only be from the molecules that were synthesised during the pulse. You're not going to accidentally label new molecules as you go along.
Then, you let time do its thing. The cell is a bustling metropolis, with molecules constantly being synthesized, modified, transported, degraded, and secreted. Your labelled molecules are now on their own adventure, participating in these cellular processes. They might move from the cytoplasm to the nucleus, get folded into a specific shape, attach to other molecules, or even be destined for the cellular trash bin.
The "chase" period is divided into multiple time points. You don't just check on them once. No, no. You take samples at various intervals – say, 15 minutes, 30 minutes, 1 hour, 2 hours, 12 hours, 24 hours. Each time point is like taking a snapshot of what’s happening to your labelled molecules at that specific moment in their journey. At each of these time points, you'll analyze your samples to see where the label has gone and how its distribution has changed.
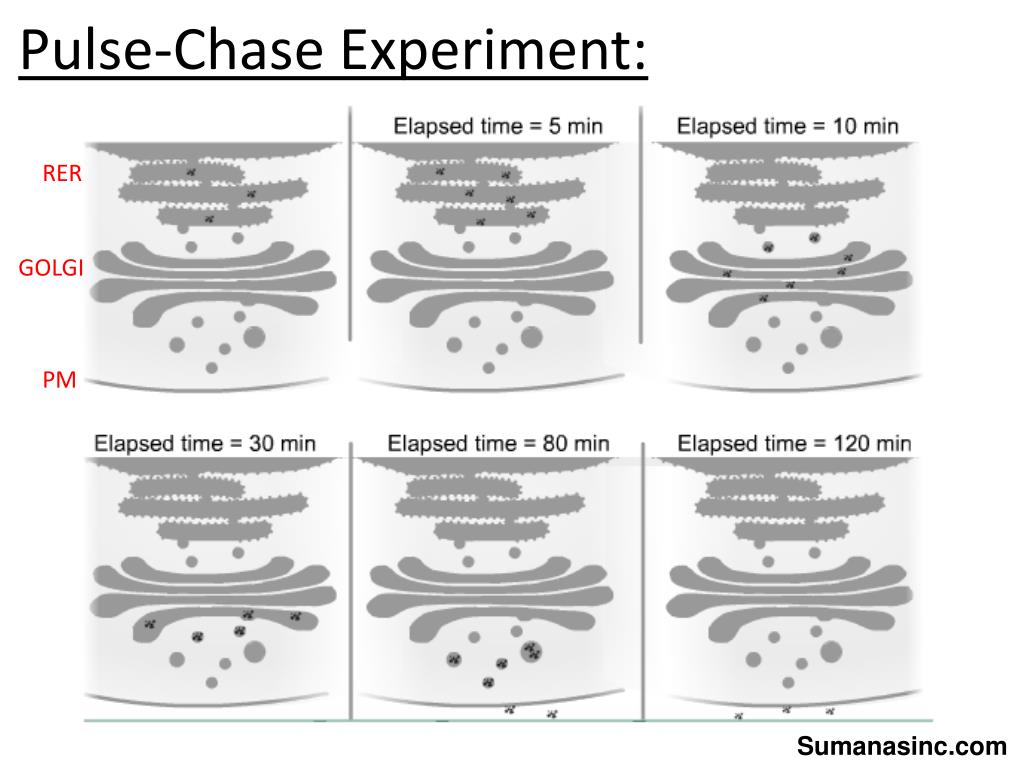
This allows you to build a narrative. You can see, for instance, if the label first appears in the endoplasmic reticulum, then moves to the Golgi apparatus, and is eventually secreted outside the cell. Or you might observe it accumulating in a specific organelle, indicating a particular function or localization. You’re essentially mapping out the pathway and the kinetics of your molecule of interest.

The "Chasing" Process: From Snapshot to Story
Once you have your labelled molecules in hand (metaphorically speaking, of course!), the next critical step in the chase is isolating and quantifying the label at each time point. This is where you translate those snapshots into a coherent story. And let me tell you, it's a whole science in itself!
How do you actually see where the label has gone? Well, it depends on your label and your system. If you used radioactive isotopes, you might use techniques like scintillation counting to measure the amount of radioactivity. This is a direct and sensitive way to quantify how much of your labelled molecule is present. Imagine having a little Geiger counter that can tell you exactly how much "glow" is in each sample.
If you used fluorescent labels, then microscopy becomes your best friend. You’ll be using confocal microscopy, fluorescence microscopy, or even super-resolution microscopy to visualize the location of your labelled molecules within the cells. This is where you can actually see your molecule zipping around, accumulating in specific compartments, or even interacting with other cellular structures. It’s like watching a tiny, invisible movie playing out inside your cells. Pretty mind-blowing, right?
Another common technique, especially when you’re interested in proteins, is to use immunoprecipitation. You can design an antibody that specifically binds to your protein of interest. Then, you can use this antibody to pull down your labelled protein from the cell lysate. Afterwards, you measure the radioactivity or fluorescence in the immunoprecipitated complex. This allows you to specifically track your protein even if it's in a complex mixture.
The analysis of this data is where the magic truly happens. You're not just collecting numbers or pretty pictures; you're piecing together a dynamic picture of molecular trafficking and transformation. You'll be looking for trends: Does the signal decrease in one compartment and increase in another? Does the overall signal intensity change over time, indicating synthesis or degradation? Are there any unexpected accumulations or disappearances?
This is also where you have to be careful about experimental artifacts. Did you wash away all the unincorporated label properly? Is your antibody specific enough? Could the cells themselves be reacting to the labelling process in some unforeseen way? It’s a constant dance between observing the biological process and critically evaluating your experimental methodology. It requires a lot of attention to detail, a good understanding of the biology you’re studying, and a healthy dose of skepticism.
Ultimately, the "chase" is about building a kinetic profile of your molecule. It's about understanding not just where it is, but how long it takes to get there, how long it stays there, and what happens to it along the way. It's the journey from a fleeting moment of introduction (the pulse) to a comprehensive understanding of a molecule's life cycle within a living system.
Putting It All Together: The Power of the Pulse-Chase
So, to circle back to our original question: What is the first step in a pulse-chase experiment? It's undeniably the careful selection and introduction of a labelled precursor molecule. Without a proper pulse, the chase is impossible. You'd be chasing ghosts, or worse, chasing nothing at all. It's the foundational element that makes the entire intricate process of tracking molecules possible.
It’s a technique that, despite its seemingly simple premise, unlocks incredibly complex biological questions. It helps us understand protein synthesis and secretion, DNA replication and repair, RNA processing, metabolic pathways, and so much more. It's like having a temporal lens that allows us to see the dynamic, living processes within cells, rather than just static snapshots.
And the beauty of it is its versatility. You can adapt it to study almost any molecule that can be labelled and is incorporated into cellular processes. From the smallest metabolites to the largest protein complexes, the pulse-chase offers a window into their world. It's a testament to the power of clever experimental design and the ingenuity of scientists trying to unravel the mysteries of life, one labelled molecule at a time.
So next time you hear about a pulse-chase experiment, remember it’s not just about the chase. It’s about that initial, crucial moment where you give your molecules a voice, a signal, a way to tell their story. And that, my friends, is where all the scientific adventure truly begins.
