How Does Shotgun Cloning Differ From The Clone-by-clone Method

So, you’ve probably heard the buzz about cloning. It’s not just for sheep anymore, right? Scientists are getting really creative with how they copy DNA. And when we talk about copying DNA, two cool methods pop into the spotlight: shotgun cloning and clone-by-clone. They sound kinda similar, but trust me, they’re like chalk and cheese. Or maybe more like a super-organized librarian versus a party-throwing DJ.
Let’s dive in and see what makes them tick. It’s actually pretty fascinating, and dare I say, a little bit fun to think about!
The DJ Approach: Shotgun Cloning
Imagine you have a massive library, like, all the books in the world. And you need to find a specific sentence in one of those books. How would you do it?
Well, shotgun cloning is like saying, "Forget finding the exact book and page! Let's just chop up all the books into tiny, tiny pieces. Then we'll mix all those pieces up, and see if we can somehow put them back together to find our sentence. And maybe even read the whole paragraph!"
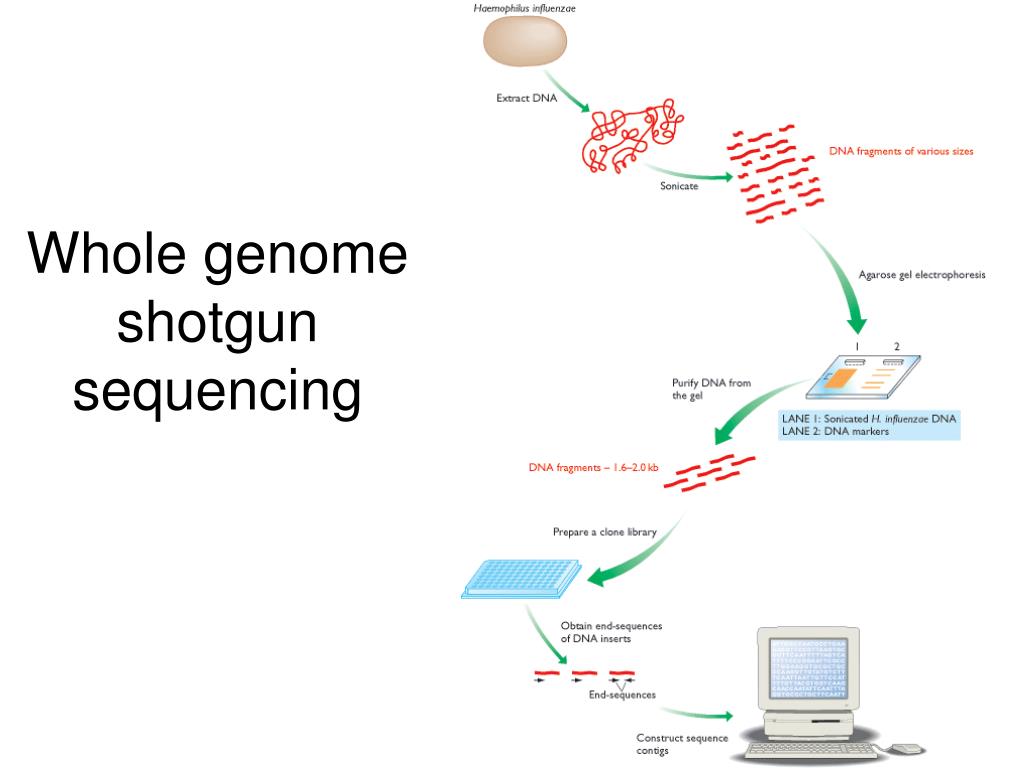
That’s the essence of shotgun cloning. Instead of carefully picking out one gene or section at a time, scientists take the entire genome – the complete set of DNA in an organism – and blast it into millions of tiny fragments. Think of it as a genetic confetti cannon!
These fragments are then inserted into special carriers, usually something called a plasmid. Think of a plasmid as a tiny, circular, genetically modified frisbee that can carry a piece of DNA. It’s like giving our DNA confetti a ride into a new host, often bacteria, where it can be copied many, many times.
Once you have tons of these little frisbees, each carrying a random snippet of the original genome, the real fun begins. Scientists then sequence the DNA on each of these fragments. So, they read the letters (A, T, C, G) of each tiny piece.
Now, this is where the magic, and a bit of computer wizardry, comes in. They have a giant pile of DNA sequences, all jumbled up. The challenge is to put them back in the right order to reconstruct the original genome. It's like solving a jigsaw puzzle where you have millions of pieces, and you don't even have the picture on the box!
Computers are the superheroes here. They look for overlapping sequences between the fragments. If one fragment ends with "ATGC" and another begins with "TGCAT", the computer knows they likely came from the same spot and can be stitched together. It’s a massive computational effort, but it’s incredibly powerful.
Why is it called "shotgun"? Because it’s like firing a shotgun into a crowd of information. You hit a lot of targets randomly, and then you have to sort through the mess to find what you’re looking for. It's a broad, aggressive strategy.
A quirky fact: The first time this was used on a large scale, for sequencing the human genome, it was revolutionary! It allowed us to tackle huge projects that would have been impossible otherwise. Imagine the patience of those early scientists!
The Librarian Approach: Clone-by-Clone
Now, let’s switch gears and think about our super-organized librarian. If they needed to find that specific sentence in the massive library, they wouldn't start by shredding every book, would they?
No, they’d probably have a catalog. They’d know which section the book is in, maybe even which shelf. They’d be methodical and targeted.
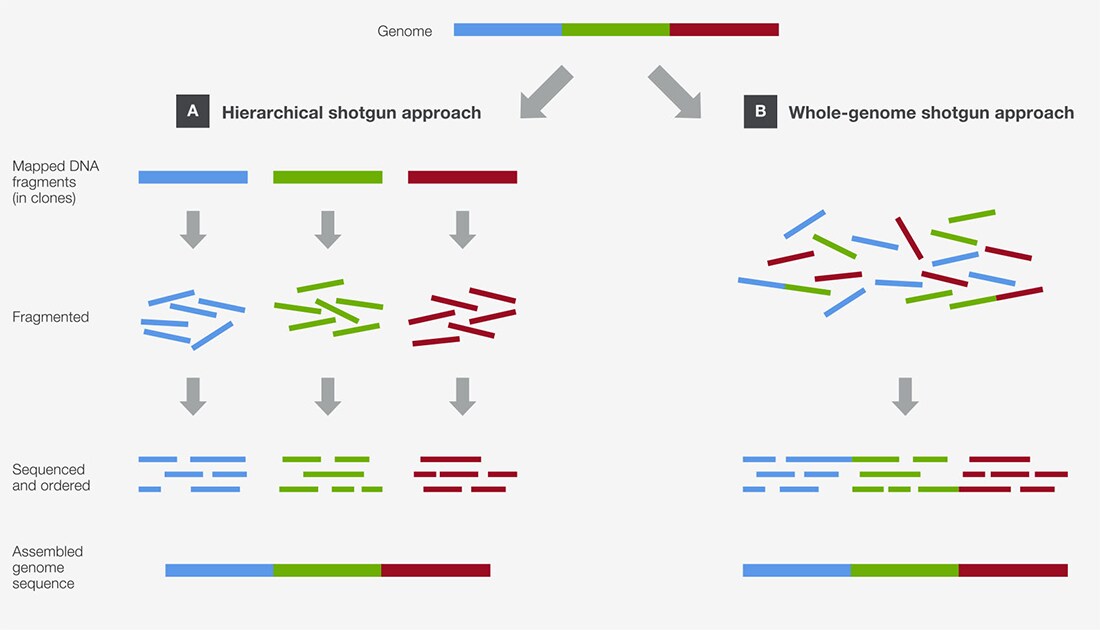
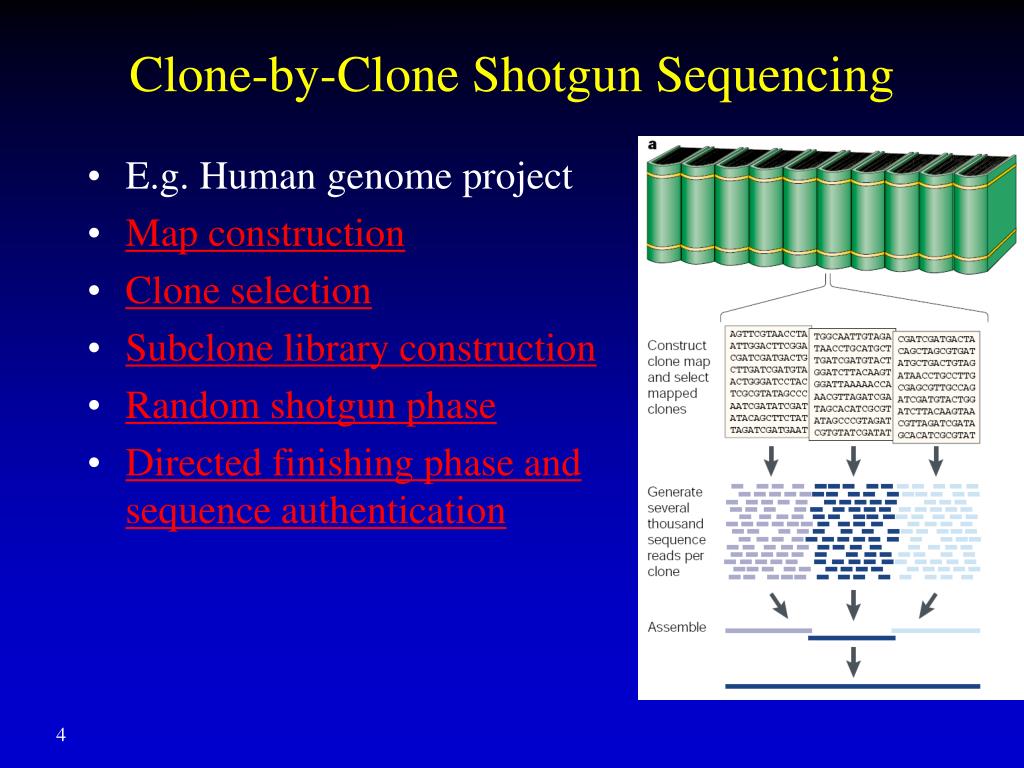
That’s exactly what clone-by-clone, also sometimes called the hierarchical shotgun sequencing, is all about. Instead of blasting the whole genome, this method breaks it down into much larger chunks first.
Think of it this way: first, the genome is cut into manageable pieces, like chapters of a book. These larger pieces are then inserted into special cloning vectors, often called BACs (Bacterial Artificial Chromosomes) or YACs (Yeast Artificial Chromosomes). These are like bigger, more robust carriers than plasmids. They’re designed to hold much longer stretches of DNA.
Once these big chunks are cloned and copied, scientists then take each individual chunk and perform shotgun sequencing on that chunk alone. So, they’ll take one BAC, chop it up into millions of tiny pieces, sequence those pieces, and then reassemble that one BAC sequence.
It's like taking a chapter from the library, chopping only that chapter into tiny pieces, and then reassembling that chapter. Then you move on to the next chapter and repeat the process.
This method provides a lot of guidance. Because you know which large chunk each set of fragments came from, reassembling the entire genome becomes much easier. It’s like having the table of contents to help you put the books back together. The computer has a much clearer roadmap.
This approach is like being a detective. You have clues (the larger chunks), and you use those clues to meticulously piece together the bigger picture. It’s a more controlled, stepwise approach.
A funny detail: While shotgun cloning is faster at generating a lot of raw sequence data, clone-by-clone can sometimes be more accurate for assembling complex genomes because it’s less prone to errors caused by repetitive DNA sequences. It’s like the librarian being super careful not to misplace a single page.

Why the Fuss?
So, why should you care about these technical terms? Because they represent different ways of tackling a massive biological puzzle. Both methods aim to read the entire DNA code of an organism.
Shotgun cloning is like a whirlwind tour. It’s fast, it’s ambitious, and it relies heavily on powerful computers to sort out the chaos. It’s great for getting a quick overview or when you have a really, really big genome to tackle.
Clone-by-clone is like a meticulously planned expedition. It takes more upfront effort to break things down, but the assembly process is often smoother and more reliable, especially for tricky genomes.
The choice between them often depends on the specific project, the organism’s genome complexity, and the available technology. It’s like choosing between a sprinter and a marathon runner – both get to the finish line, but in very different ways!
And hey, the more we understand how to read and assemble genomes, the more we unlock the secrets of life. It’s pretty cool to think that a messy shotgun blast or a librarian’s careful organization can lead to such incredible discoveries. Makes you wonder what other awesome cloning tricks are out there, doesn't it?
